The changes in the probability distribution for small increments of time are given by: Letīe the transition rate from state to state. For a fixed site, letīe the column vector of probabilities of states and at time. Assume that the processes followed by the m sites are Markovian independent, identically distributed and constant in time. Theorem: Continuous-time transition matrices satisfy:Ĭonsider a DNA sequence of fixed length m evolving in time by base replacement. This then allows us to write that probability as. Where the top-left and bottom-right blocks correspond to transition probabilities and the top-right and bottom-left blocks corresponds to transversion probabilities.Īssumption: If at some time, the Markov chain is in state, then the probability that at time, it will be in state depends only upon, and. The corresponding transition matrices will look like: Jukes-Cantor, Kimura, etc.) in a continuous time fashion. Įxample: We would like to model the substitution process in DNA sequences ( i.e. Where each individual entry, refers to the probability that state will change to state in time. 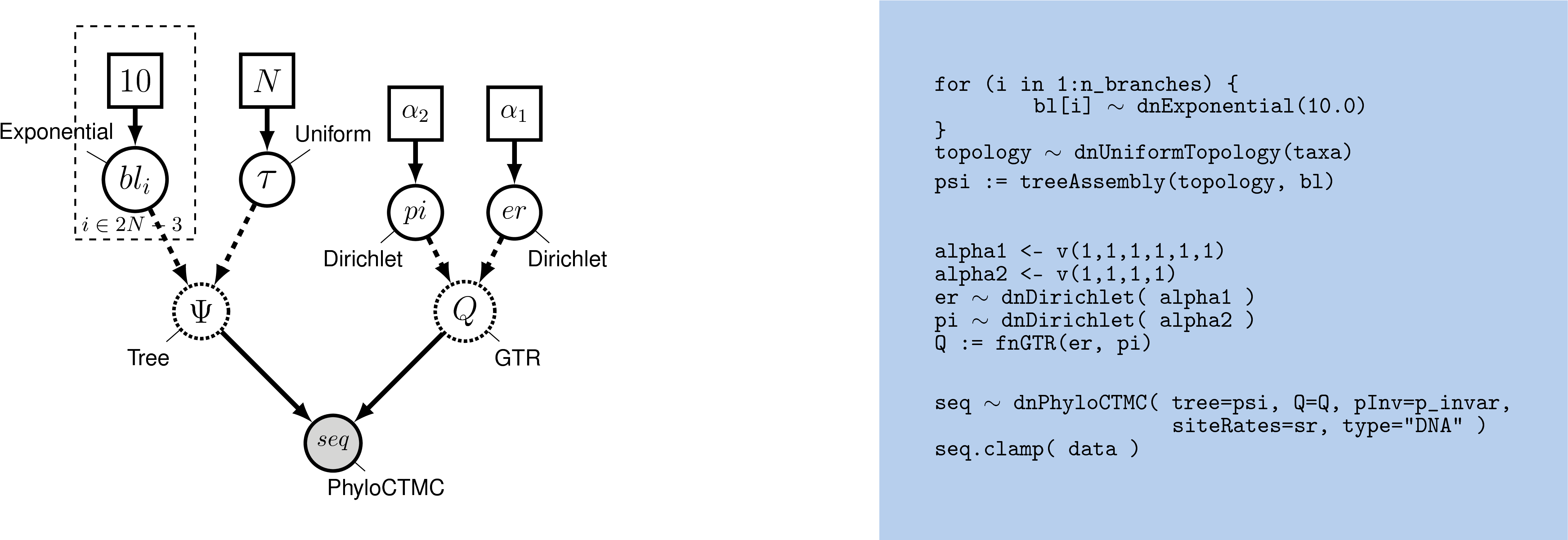
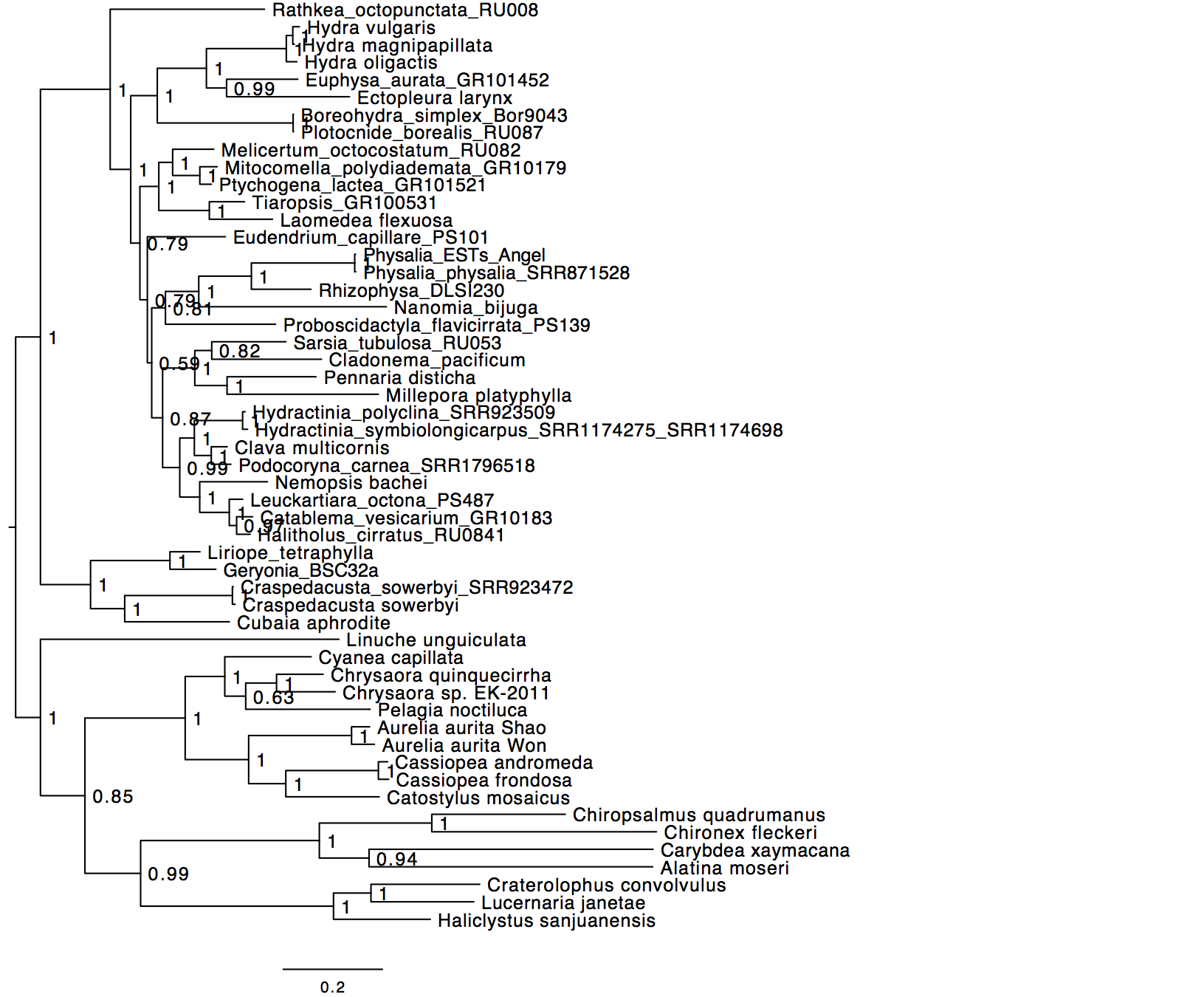
Specifically, if are the states, then the transition matrix Which are, in addition, parameterized by time. 2.4 HKY85 model (Hasegawa, Kishino and Yano 1985)ĭNA Evolution as a Continuous Time Markov Chain Continuous Time Markov ChainsĬontinuous-time Markov chains have the usual transition matrices.2.1 JC69 model (Jukes and Cantor, 1969).1.2 Deriving the Dynamics of Substitution.1 DNA Evolution as a Continuous Time Markov Chain.Therefore, the bias introduced by the use of the GTR + Γ model to analyze datasets, in which the time-reversibility and stationarity assumptions are violated, is likely not large and can be reduced by applying multiple calibrations.Īll datasets are deposited in Figshare. Confidence and credibility intervals from GTR + Γ analysis usually contained correct times. The use of only a few calibrations reduced these biases considerably (∼5%). Divergence times obtained using a GTR + Γ model differed only slightly (∼3% on average) from the expected times for NR datasets, but the difference was larger for NS datasets (∼10% on average). We tested Bayesian and RelTime approaches that do not require a molecular clock for estimating divergence times. We quantified the bias on time estimates that resulted from using the GTR + Γ model for the analysis of computer-simulated nucleotide sequence alignments that were evolved with non-stationary (NS) and non-reversible (NR) substitution models. Many reports have quantified the impact of violations of these underlying assumptions on molecular phylogeny, but none have systematically analyzed their impact on divergence time estimates. Frequently the same General Time Reversible (GTR) model across lineages along with a gamma (+Γ) distributed rates across sites is used in relaxed clock analyses, which assumes time-reversibility and stationarity of the substitution process. No software tools for molecular dating allow researchers to relax these two assumptions in their data analyses. As the number and diversity of species and genes grow in contemporary datasets, two common assumptions made in all molecular dating methods, namely the time-reversibility and stationarity of the substitution process, become untenable. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
